192 JOURNAL OF COSMETIC SCIENCE TREATMENT OF NHEKS WITH AGE-BSA NHEKs were preliminarily cultured in an EpiLife® medium in the presence or absence of an agent (bafilomycin A1, chloroquine, EX527, SRT1720, resveratrol, or NMN). After preincubation, AGE-BSA was added to the NHEKs, and the cells were further cultured for 24 hours. After incubation, the cells were subjected to intracellular NAD+ measurement, immunoblot analysis, or real-time polymerase chain reaction (PCR). For UV irradiation tests, after a 24-hour preliminary incubation, the cells were exposed to UVA/B in a phenol red-free EpiLife® medium (Thermo Fisher Scientific). After exposure to UV, the cells were cultured in the presence or absence of AGE-BSA in an EpiLife® medium for the indicated time periods. After incubation, the cells were subjected to immunoblot analysis or real-time polymerase chain reaction (PCR). UVA/B EXPOSURE NHEKs in the EpiLife® culture media without phenol red were exposed to UVA/B (wavelength 280–400 nm UVA: 0, 6, 13, and 19 J/cm2 and UVB: 0, 12, 23, and 35 mJ/cm2) at room temperature using a xenon arc lamp-based sunlight simulator (Wacom, Saitama, Japan). Calibration of the UVA/B dosage was performed using the MS-211-I UV dosimeter (EKO, Tokyo, Japan). QUANTIFICATION OF RESIDUAL AGE-BSA IN THE MEDIUM NHEKs were preliminarily cultured in the EpiLife® medium. After 24 hours of incubation, the cells were exposed to UVA/B and cultured with AGE-BSA in the phenol red-free EpiLife® medium for the indicated time periods. The culture medium was applied to quantify residual AGE-BSA in the medium that was not taken up by the cells. Briefly, we measured the characteristic fluorescence of AGE-BSA (excitation wavelength of 370 nm and emission wavelength of 440 nm) using an Infinite 200 Pro plate reader (Tecan, Männedorf, Switzerland). QUANTIFICATION OF NAD+ CONTENT The amount of NAD+ was measured using an NAD+/NADH Assay Kit-WST (Dojindo Laboratories, Kumamoto, Japan) according to the manufacturer’s instructions. A sample solution of cellular NAD+ was prepared using the attached lysis buffer and purified using a 10-kDa molecular weight cutoff device (Merck Millipore). Each sample was used for the subsequent enzymatic reaction. The absorbance of the samples at 450 nm was measured using an Infinite 200 Pro plate reader (Tecan). The NAD+ levels were standardized to their respective protein concentrations calculated by BCA protein quantification. SODIUM DODECYL SULFATE−POLYACRYLAMIDE GEL ELECTROPHORESIS (SDS−PAGE) AND IMMUNOBLOT ANALYSIS Cell lysates were prepared with a cold lysis buffer (50 mM Tris, pH 8.0, 150 mM NaCl, 0.5% sodium deoxycholate, 0.1% SDS, 1% NP-40) containing a 1 × protease inhibitor
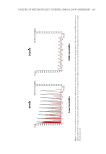
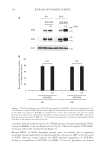
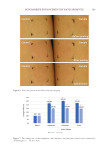
193 Accumulation of Advanced Glycation End Products cocktail (Hoffmann-La Roche, Basel, Switzerland). The proteins were run on 8% SDS- polyacrylamide gels, were transferred to polyvinylidene fluoride membranes, and were incubated with 5% skim milk in a Tris-buffered saline containing 0.1% Tween-20 at room temperature for 1 hour. After blocking, the membranes were treated overnight with the primary antibody at 4°C. After washing, the blots were further incubated for 1 hour at room temperature with the HRP-conjugated IgG antibody. The ECL Prime Western Blotting Detection Reagent (Cytiva, Marlborough, MA, USA) and LAS-3000 mini (Fujifilm, Tokyo, Japan) were used for chemiluminescence detection. REAL-TIME PCR After NHEKs were treated as indicated, total RNA was collected using the RNeasy Mini RNA isolation kit (Qiagen, Mississauga, Canada). RNA was eluted into RNase-free water and aliquoted at −80°C until use. Reverse transcription-PCR was performed on 1 µg of total RNA using a PrimeScript II 1st strand cDNA Synthesis Kit (Takara Bio, Shiga, Japan). Real-time PCR was performed using PrimeTime® Std qPCR Assay (Integrated DNA Technologies, Coralville, Iowa, USA) for glyoxalase 1 (GLO1 Hs.PT.58.20622948), glyoxalase 2 (GLO2 Hs.PT.58.41046034), and 18S (Hs.PT.39a.22214856.g), according to the manufacturer’s instructions on the Applied Biosystems 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). 18S was used as the housekeeping gene for normalization. The comparative C T method was used to calculate the relative amount of mRNA. STATISTICAL ANALYSIS Data were expressed as means ± SEMs (standard errors of the means) from at least three independent experiments. The data were analyzed by one-way ANOVA. We performed the Tukey–Kramer test to analyze the differences among the means. The statistical significances were set at p 0.05 and p 0.01. Asterisks in the illustrations herein indicate statistical significances compared to the control (*p 0.05 and **p 0.01). RESULTS We first investigated if UV irradiation promotes AGE accumulation in epidermal keratinocytes. After the NHEKs were exposed to UVA/B irradiation (280–400 nm), AGE- BSA was added to them as exogenous AGEs. Using immunoblotting with anti-AGE and anti-BSA antibodies, we confirmed that the added AGE-BSA was incorporated into and accumulated in NHEKs (Figure 1A). The intracellular AGE-BSA content significantly increased 8 and 24 hours after UVA/B irradiation. This result suggests that UV exposure enhances the accumulation of exogenous AGEs in the cells. The residual AGE-BSA levels in the medium decreased by 20–30% after 8 and 24 hours of incubation, whereas UVA/B irradiation did not cause any further reduction in residual levels of AGEs in the culture medium. This suggests that UV radiation did not specifically promote the uptake of exogenous AGEs by the cells (Figure 1B). In addition, the autophagy inhibitors bafilomycin A1 and chloroquine increased subcellular AGE-BSA levels (Figure 2). These results indicate that exogenous AGEs incorporated into NHEKs could be degraded via autophagy. In
Purchased for the exclusive use of nofirst nolast (unknown) From: SCC Media Library & Resource Center (library.scconline.org)