634 JOURNAL OF COSMETIC SCIENCE
microbiome.2 Understanding these interactions is crucial for deciphering their impact on
health and disease, paving the way for targeted interventions and therapies that harness the
microbiome’s potential to support overall well-being.
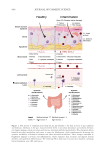
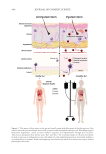
Furthermore, the interactions between the gut microbiome and the skin, commonly referred
to as the “gut-skin axis,” have been increasingly clarified. Research has firmly established
that the gut-skin axis represents a pathway of communication between the gastrointestinal
(GI) tract and the skin, highlighting the interconnectedness of these two organ systems.3
While the terms “skin-gut axis” and “gut-skin axis” are often used interchangeably in
research, this review will specifically use “gut-skin axis” to denote the communication
pathway from the gut microbiome to the skin. The mechanisms underlying this link
from the gut microbiome to the skin include immunomodulation, wherein the gut
microbiome influences the systemic immune response, impacting skin inflammation and
immune function.4 Hormonal pathways also play a crucial role, as hormones produced in
response to gut microbial activity can affect skin physiology.3 Additionally, the production
of metabolites, such as short-chain fatty acids (SCFAs) by gut bacteria, can enter the
bloodstream and exert effects on skin cells, contributing to the regulation of skin barrier
function and inflammatory processes.5 Interestingly, emerging research has postulated the
bidirectional nature of this axis, suggesting that the state of the skin may, in turn, influence
the gut microbiome, known in this review as the “skin-gut axis.”6
Historically, microbiome studies primarily relied on culture-based techniques to identify
microorganisms present in samples.2Whilevaluableatthetime,thesemethodshadsignificant
limitations, including an inability to accurately represent the endogenous microbiome
environment and difficulties in culturing many microorganisms.7 Therefore, the introduction
of DNA-based culture-independent molecular techniques in the 1980s revolutionized
microbiome research.7 This paradigm shift involving analyzing the microbial DNA directly
from samples, instead of DNA from lab-grown cultures, redirected researchers’ focus toward
understanding microbial functions and interactions. Hence, metagenomics, which refers to
the study of the genetic material of entire communities of organisms, has emerged as a
pivotal technique.7 Ultimately, the “multi-omics” approach—comprising metagenomics and
additionally, metatranscriptomics, proteomics, lipidomics, and metabolomics—has become
standard practice for microbiome studies and these comprehensive methodologies are crucial
for advancing the understanding of the microbiome (summarized in Table I).
THE MULTI-OMICS APPROACH
Metagenomics is the unbiased study of the sequences and functions of all genetic information
extracted from a specific environment, such as the human skin microbiome. By employing
diverse bioinformatic systems and genomic technologies, metagenomics enables researchers
to explore the entire genome of various environmental communities and gives insight into
the diversity, structure and functional potential thereof.8 This approach was first introduced
in 1998 in reference to shotgun metagenomic sequencing. Nowadays, it is largely applied
to studies of marker genes, such as the 16S ribosomal RNA (rRNA) gene in the form
of targeted bacterial profiling (summarized in Table II).9,10 This method of microbiome
sequencing targets the 16S rRNA gene, which is highly conserved among bacteria and
archaea, although it contains hypervariable regions that can distinguish between different
species.11 Similarly, internal transcribed spacer (ITS) region amplification focuses on the
ITS region of the rRNA gene cluster, which is highly variable among different fungal
microbiome.2 Understanding these interactions is crucial for deciphering their impact on
health and disease, paving the way for targeted interventions and therapies that harness the
microbiome’s potential to support overall well-being.
Furthermore, the interactions between the gut microbiome and the skin, commonly referred
to as the “gut-skin axis,” have been increasingly clarified. Research has firmly established
that the gut-skin axis represents a pathway of communication between the gastrointestinal
(GI) tract and the skin, highlighting the interconnectedness of these two organ systems.3
While the terms “skin-gut axis” and “gut-skin axis” are often used interchangeably in
research, this review will specifically use “gut-skin axis” to denote the communication
pathway from the gut microbiome to the skin. The mechanisms underlying this link
from the gut microbiome to the skin include immunomodulation, wherein the gut
microbiome influences the systemic immune response, impacting skin inflammation and
immune function.4 Hormonal pathways also play a crucial role, as hormones produced in
response to gut microbial activity can affect skin physiology.3 Additionally, the production
of metabolites, such as short-chain fatty acids (SCFAs) by gut bacteria, can enter the
bloodstream and exert effects on skin cells, contributing to the regulation of skin barrier
function and inflammatory processes.5 Interestingly, emerging research has postulated the
bidirectional nature of this axis, suggesting that the state of the skin may, in turn, influence
the gut microbiome, known in this review as the “skin-gut axis.”6
Historically, microbiome studies primarily relied on culture-based techniques to identify
microorganisms present in samples.2Whilevaluableatthetime,thesemethodshadsignificant
limitations, including an inability to accurately represent the endogenous microbiome
environment and difficulties in culturing many microorganisms.7 Therefore, the introduction
of DNA-based culture-independent molecular techniques in the 1980s revolutionized
microbiome research.7 This paradigm shift involving analyzing the microbial DNA directly
from samples, instead of DNA from lab-grown cultures, redirected researchers’ focus toward
understanding microbial functions and interactions. Hence, metagenomics, which refers to
the study of the genetic material of entire communities of organisms, has emerged as a
pivotal technique.7 Ultimately, the “multi-omics” approach—comprising metagenomics and
additionally, metatranscriptomics, proteomics, lipidomics, and metabolomics—has become
standard practice for microbiome studies and these comprehensive methodologies are crucial
for advancing the understanding of the microbiome (summarized in Table I).
THE MULTI-OMICS APPROACH
Metagenomics is the unbiased study of the sequences and functions of all genetic information
extracted from a specific environment, such as the human skin microbiome. By employing
diverse bioinformatic systems and genomic technologies, metagenomics enables researchers
to explore the entire genome of various environmental communities and gives insight into
the diversity, structure and functional potential thereof.8 This approach was first introduced
in 1998 in reference to shotgun metagenomic sequencing. Nowadays, it is largely applied
to studies of marker genes, such as the 16S ribosomal RNA (rRNA) gene in the form
of targeted bacterial profiling (summarized in Table II).9,10 This method of microbiome
sequencing targets the 16S rRNA gene, which is highly conserved among bacteria and
archaea, although it contains hypervariable regions that can distinguish between different
species.11 Similarly, internal transcribed spacer (ITS) region amplification focuses on the
ITS region of the rRNA gene cluster, which is highly variable among different fungal