549 Modern Skincare
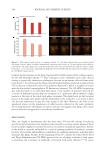
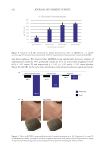
region of interest by a Canfield research device under UV-fluorescence or red-light modalities.
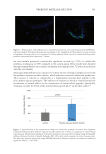
Finally, the three participants had their skin swabbed (5 samples per participant), which was
tested for relative levels of skin bacterial groups via 16S rRNA sequencing.61
REFERENCES
(1) Grice EA, Segre JA. The skin microbiome. Nat Rev Microbiol. 2011 9(4):244–253. doi:10.1038/
nrmicro2537
(2) Fournière M, Latire T, Souak D, Feuilloley MGJ, Bedoux G. Staphylococcus epidermidis and Cutibacterium
acnes: two Major Sentinels of Skin microbiota and the Influence of Cosmetics. Microorganisms. 2020 8(11).
doi:10.3390/microorganisms8111752
(3) Brüggemann H, Salar-Vidal L, Gollnick HPM, Lood R. A Janus-faced Bacterium: Host-Beneficial
and -Detrimental Roles of Cutibacterium acnes. Front Microbiol. 2021 12:673845. doi:10.3389/
fmicb.2021.673845
(4) Zheng Y, Hunt RL, Villaruz AE, et al. Commensal Staphylococcus epidermidis contributes to skin
barrier homeostasis by generating protective ceramides. Cell Host Microbe. 2022 30(3):301–313.e9.
doi:10.1016/j.chom.2022.01.004
(5) Grice EA, Kong HH, Conlan S, et al. Topographical and temporal diversity of the human skin
microbiome. Science. 2009 324(5931):1190–1192. doi:10.1126/science.1171700
(6) Oh J, Byrd AL, Park M, NISC Comparative Sequencing Program, Kong HH, Segre JA. Temporal
stability of the human skin microbiome. Cell. 2016 165(4):854–866. doi:10.1016/j.cell.2016.04.008
(7) Coenye T, Peeters E, Nelis HJ. Biofilm formation by Propionibacterium acnes is associated with increased
resistance to antimicrobial agents and increased production of putative virulence factors. Res Microbiol.
2007 158(4):386–392. doi:10.1016/j.resmic.2007.02.001
(8) Williams MR, Gallo RL. The role of the skin microbiome in atopic dermatitis. Curr Allergy Asthma Rep.
2015 15(11):65. doi:10.1007/s11882-015-0567-4
(9) Koh LF, Ong RY, Common JE. Skin microbiome of atopic dermatitis. Allergol Int. 2022 71(1):31–39.
doi:10.1016/j.alit.2021.11.001
(10) Williams HC, Dellavalle RP, Garner S. Acne vulgaris. Lancet. 2012 379(9813):361–372. doi:10.1016/
S0140-6736(11)60321-8
(11) Fitz-Gibbon S, Tomida S, Chiu B-H, et al. Propionibacterium acnes strain populations in the human
skin microbiome associated with acne. J Invest Dermatol. 2013 133(9):2152–2160. doi:10.1038/jid.2013.21
(12) Kinney MA, Yentzer BA, Fleischer AB, Feldman SR. Trends in the treatment of acne vulgaris: are
measures being taken to avoid antimicrobial resistance? J Drugs Dermatol. 2010 9(5):519–524.
(13) Uhlenhake E, Yentzer BA, Feldman SR. Acne vulgaris and depression: a retrospective examination. J
Cosmet Dermatol. 2010 9(1):59–63. doi:10.1111/j.1473-2165.2010.00478.x
(14) Yentzer BA, Hick J, Reese EL, Uhas A, Feldman SR, Balkrishnan R. Acne vulgaris in the United States:
a descriptive epidemiology. Cutis. 2010 86(2):94–99.
(15) Li ZJ, Choi DK, Sohn KC, et al. Propionibacterium acnes activates the NLRP3 inflammasome in human
sebocytes. J Invest Dermatol. 2014 134(11):2747–2756. doi:10.1038/jid.2014.221
(16) Woese CR, Fox GE. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc Natl
Acad Sci U S A. 1977 74(11):5088–5090. doi:10.1073/pnas.74.11.5088
(17) Grice EA. The skin microbiome: potential for novel diagnostic and therapeutic approaches to cutaneous
disease. Semin Cutan Med Surg. 2014 33(2):98–103. doi:10.12788/j.sder.0087
(18) Grice EA. The intersection of microbiome and host at the skin interface: genomic- and metagenomic-
based insights. Genome Res. 2015 25(10):1514–1520. doi:10.1101/gr.191320.115
(19) Jamal M, Bukhari SMAUS, Andleeb S, et al. Bacteriophages: an overview of the control strategies against
multiple bacterial infections in different fields. J Basic Microbiol. 2019 59(2):123–133. doi:10.1002/
jobm.201800412
region of interest by a Canfield research device under UV-fluorescence or red-light modalities.
Finally, the three participants had their skin swabbed (5 samples per participant), which was
tested for relative levels of skin bacterial groups via 16S rRNA sequencing.61
REFERENCES
(1) Grice EA, Segre JA. The skin microbiome. Nat Rev Microbiol. 2011 9(4):244–253. doi:10.1038/
nrmicro2537
(2) Fournière M, Latire T, Souak D, Feuilloley MGJ, Bedoux G. Staphylococcus epidermidis and Cutibacterium
acnes: two Major Sentinels of Skin microbiota and the Influence of Cosmetics. Microorganisms. 2020 8(11).
doi:10.3390/microorganisms8111752
(3) Brüggemann H, Salar-Vidal L, Gollnick HPM, Lood R. A Janus-faced Bacterium: Host-Beneficial
and -Detrimental Roles of Cutibacterium acnes. Front Microbiol. 2021 12:673845. doi:10.3389/
fmicb.2021.673845
(4) Zheng Y, Hunt RL, Villaruz AE, et al. Commensal Staphylococcus epidermidis contributes to skin
barrier homeostasis by generating protective ceramides. Cell Host Microbe. 2022 30(3):301–313.e9.
doi:10.1016/j.chom.2022.01.004
(5) Grice EA, Kong HH, Conlan S, et al. Topographical and temporal diversity of the human skin
microbiome. Science. 2009 324(5931):1190–1192. doi:10.1126/science.1171700
(6) Oh J, Byrd AL, Park M, NISC Comparative Sequencing Program, Kong HH, Segre JA. Temporal
stability of the human skin microbiome. Cell. 2016 165(4):854–866. doi:10.1016/j.cell.2016.04.008
(7) Coenye T, Peeters E, Nelis HJ. Biofilm formation by Propionibacterium acnes is associated with increased
resistance to antimicrobial agents and increased production of putative virulence factors. Res Microbiol.
2007 158(4):386–392. doi:10.1016/j.resmic.2007.02.001
(8) Williams MR, Gallo RL. The role of the skin microbiome in atopic dermatitis. Curr Allergy Asthma Rep.
2015 15(11):65. doi:10.1007/s11882-015-0567-4
(9) Koh LF, Ong RY, Common JE. Skin microbiome of atopic dermatitis. Allergol Int. 2022 71(1):31–39.
doi:10.1016/j.alit.2021.11.001
(10) Williams HC, Dellavalle RP, Garner S. Acne vulgaris. Lancet. 2012 379(9813):361–372. doi:10.1016/
S0140-6736(11)60321-8
(11) Fitz-Gibbon S, Tomida S, Chiu B-H, et al. Propionibacterium acnes strain populations in the human
skin microbiome associated with acne. J Invest Dermatol. 2013 133(9):2152–2160. doi:10.1038/jid.2013.21
(12) Kinney MA, Yentzer BA, Fleischer AB, Feldman SR. Trends in the treatment of acne vulgaris: are
measures being taken to avoid antimicrobial resistance? J Drugs Dermatol. 2010 9(5):519–524.
(13) Uhlenhake E, Yentzer BA, Feldman SR. Acne vulgaris and depression: a retrospective examination. J
Cosmet Dermatol. 2010 9(1):59–63. doi:10.1111/j.1473-2165.2010.00478.x
(14) Yentzer BA, Hick J, Reese EL, Uhas A, Feldman SR, Balkrishnan R. Acne vulgaris in the United States:
a descriptive epidemiology. Cutis. 2010 86(2):94–99.
(15) Li ZJ, Choi DK, Sohn KC, et al. Propionibacterium acnes activates the NLRP3 inflammasome in human
sebocytes. J Invest Dermatol. 2014 134(11):2747–2756. doi:10.1038/jid.2014.221
(16) Woese CR, Fox GE. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc Natl
Acad Sci U S A. 1977 74(11):5088–5090. doi:10.1073/pnas.74.11.5088
(17) Grice EA. The skin microbiome: potential for novel diagnostic and therapeutic approaches to cutaneous
disease. Semin Cutan Med Surg. 2014 33(2):98–103. doi:10.12788/j.sder.0087
(18) Grice EA. The intersection of microbiome and host at the skin interface: genomic- and metagenomic-
based insights. Genome Res. 2015 25(10):1514–1520. doi:10.1101/gr.191320.115
(19) Jamal M, Bukhari SMAUS, Andleeb S, et al. Bacteriophages: an overview of the control strategies against
multiple bacterial infections in different fields. J Basic Microbiol. 2019 59(2):123–133. doi:10.1002/
jobm.201800412