588 JOURNAL OF COSMETIC SCIENCE
hours the Cutibacterium species were seeded at 106–108 CFU/mL in an insert above the
keratinocyte’s cultures grown in KSFM medium for 48 hours. To evaluate the impact
of the different C. acnes strains on the cell viability, a MTT assay was performed. The
MTT assay is a rapid colorimetric assay based on the cleavage of the tetrazolium ring of
MTT (3-(4,5-dimethylthazolk-2-yl)-2,5-diphenyl tetrazolium bromide) by dehydrogenases
in active mitochondria of living cells as an estimate of viable cell number. Sodium dodecyl
sulfate at 0.025% was used as cytotoxic positive control.
To study direct interaction, normal human keratinocytes were seeded in 96-well culture
plates at 50,000 cells/cm2 in KSFM (Thermo Fisher Scientific). C. acnes strains were heat-
killed at 95°C for 1 hour with a heated dry bath and were then seeded at 106–108 CFU/
mL in direct interaction with keratinocytes for 24 hours. Heat-killed C. acnes strains were
used instead of living C. acnes strains to avoid their interaction in MTT viability test run
in parallel to normalize the IL-8 secretion from keratinocytes. To evaluate the impact of
the different C. acnes strains on the secretion of IL-8 by keratinocytes, the media was then
collected and assessed using an IL-8 immunoassay, a technique relying on the ability of an
antibody to recognize and bind a specific macromolecule (IL-8). An inflammatory cocktail
(12.5 ng/ml of tumor necrosis factor alpha (TNFα), 12.5 ng/mL of interferon gamma
(IFNγ), and 0.5 µg/mL of Poly I:C) was used as positive control. Statistics: n =3, Student
t-test or Anova versus Untreated, *p 0.05, ***p 0.001, NS: not significant.
RESULTS
THE MICROBIAL SEQUENCING ANALYSIS BETWEEN NORMAL AND SENSITIVE SKIN
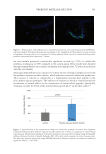
The analysis was completed for 73 volunteers (40 NS subjects and 33 SS subjects). The
diversity was analyzed using the Shannon index and highlighted no significant differences
between the two cohorts (Figure 1).
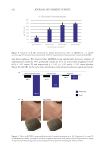
The prevalence of the species was analyzed by determining the absence or presence
of different bacterial genera on NS or SS. Figure 2 represents the 20 genera having a
Figure 1. The alpha diversity analysis between normal skin and sensitive skin by using the Shannon index.
hours the Cutibacterium species were seeded at 106–108 CFU/mL in an insert above the
keratinocyte’s cultures grown in KSFM medium for 48 hours. To evaluate the impact
of the different C. acnes strains on the cell viability, a MTT assay was performed. The
MTT assay is a rapid colorimetric assay based on the cleavage of the tetrazolium ring of
MTT (3-(4,5-dimethylthazolk-2-yl)-2,5-diphenyl tetrazolium bromide) by dehydrogenases
in active mitochondria of living cells as an estimate of viable cell number. Sodium dodecyl
sulfate at 0.025% was used as cytotoxic positive control.
To study direct interaction, normal human keratinocytes were seeded in 96-well culture
plates at 50,000 cells/cm2 in KSFM (Thermo Fisher Scientific). C. acnes strains were heat-
killed at 95°C for 1 hour with a heated dry bath and were then seeded at 106–108 CFU/
mL in direct interaction with keratinocytes for 24 hours. Heat-killed C. acnes strains were
used instead of living C. acnes strains to avoid their interaction in MTT viability test run
in parallel to normalize the IL-8 secretion from keratinocytes. To evaluate the impact of
the different C. acnes strains on the secretion of IL-8 by keratinocytes, the media was then
collected and assessed using an IL-8 immunoassay, a technique relying on the ability of an
antibody to recognize and bind a specific macromolecule (IL-8). An inflammatory cocktail
(12.5 ng/ml of tumor necrosis factor alpha (TNFα), 12.5 ng/mL of interferon gamma
(IFNγ), and 0.5 µg/mL of Poly I:C) was used as positive control. Statistics: n =3, Student
t-test or Anova versus Untreated, *p 0.05, ***p 0.001, NS: not significant.
RESULTS
THE MICROBIAL SEQUENCING ANALYSIS BETWEEN NORMAL AND SENSITIVE SKIN
The analysis was completed for 73 volunteers (40 NS subjects and 33 SS subjects). The
diversity was analyzed using the Shannon index and highlighted no significant differences
between the two cohorts (Figure 1).
The prevalence of the species was analyzed by determining the absence or presence
of different bacterial genera on NS or SS. Figure 2 represents the 20 genera having a
Figure 1. The alpha diversity analysis between normal skin and sensitive skin by using the Shannon index.